
The largest strength of mNGS is that it is an unbiased hypothesis-free diagnostic method, unlike targeted polymerase chain reaction (PCR) methods that rely on primers for identification of specific targets to be amplified and detected. What Are the Benefits of Metagenomic Next Generation Sequencing? These contigs are aligned to a reference database for taxonomic classification. (5) Contigs of long DNA stretches are assembled from shorter, overlapping sequences. (4) Human-related DNA sequence reads are removed. (3) The fragments of DNA are simultaneously and independently sequenced. 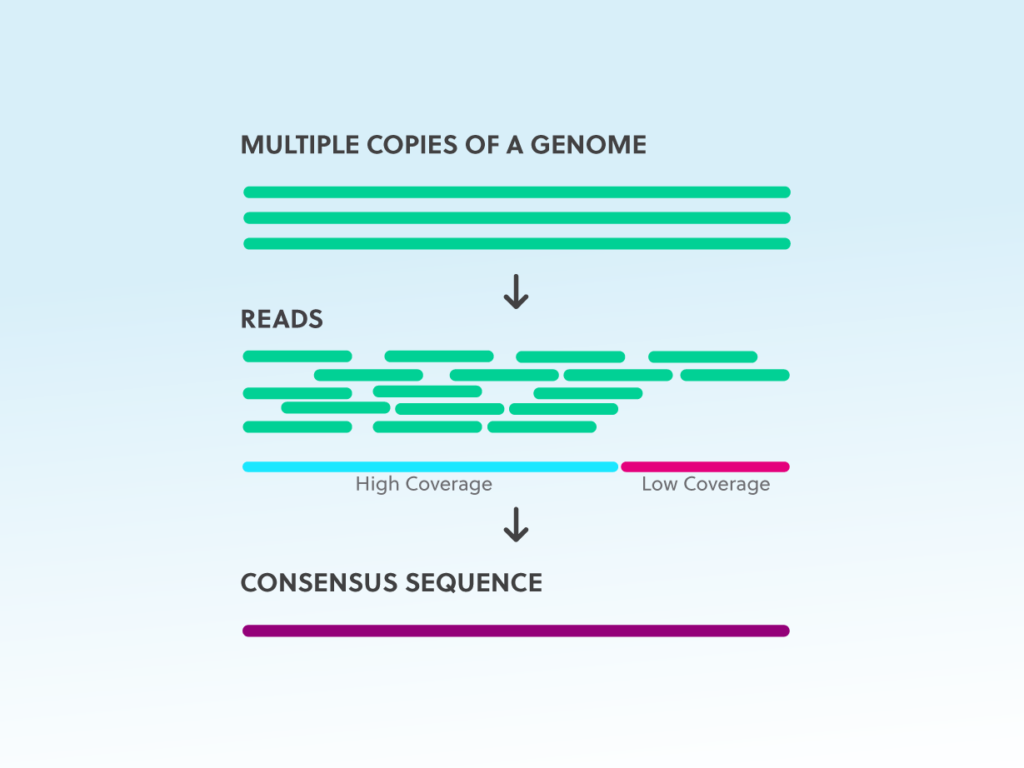
(2) Adaptors are attached for barcoding and library sequencing preparation. (1) Genomic DNA is extracted and fragmented. Workflow for metagenomic next-generation sequencing. mNGS, sometimes called shotgun sequencing, of clinical samples has been applied to various sample types including cerebrospinal fluid, blood, respiratory samples, gastrointestinal fluid, and ocular fluid. The possible clinical applications are tremendous, including diagnosis of infectious diseases, outbreak tracking, infection control surveillance, and mutation and pathogen discovery, among many others. The ability to sequence and identify nucleic acids from multiple different taxa for metagenomic analysis makes this a powerful new platform that can simultaneously identify genetic material from entirely different kingdoms of organisms. Metagenomic NGS (mNGS) is simply running all nucleic acids in a sample, which may contain mixed populations of microorganisms, and assigning these to their reference genomes to understand which microbes are present and in what proportions. This technology represents a substantial advance over the early days of sequencing when a single bacterial genome project could take several years. The constructed genome (likely containing gaps) is aligned to a reference database for identification of the organism. There are often gaps between contigs and although high-fidelity longer sequence reads would be the ideal method of sequencing, platforms that produce shorter reads are generally less costly and the overlap in sequences makes them more accurate. Several overlapping sequence reads are pieced together to produce a single longer sequence called a contig. The sequences are assembled into a single genome using computational approaches. To characterize a bacterial genome using NGS, for example, the genome is split into multiple fragments that produce sequences or reads ranging from hundreds to tens of thousands of bases in length. Contrast this technique to classical methods such as Sanger sequencing (also known as dideoxynucleotide chain termination sequencing), which processes one nucleotide sequence per reaction. Next generation sequencing is any of several high-throughput sequencing methods whereby billions of nucleic acid fragments can be simultaneously and independently sequenced. What exactly is mNGS and how is it different from the many other nucleic acid technologies out there? What Is Metagenomic Next Generation Sequencing? A deep dive point-counterpoint discussion from the Journal of Clinical Microbiology discusses the challenges and opportunities that may come with introduction of metagenomic next generation sequencing (mNGS) into routine laboratories. There is nonetheless considerable controversy on how, when, and where next generation sequencing will play a role in the clinical diagnostic laboratory. Next generation sequencing (NGS) methods started to appear in the literature in the mid-2000s and had a transformative effect on our understanding of microbial genomics and infectious diseases.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
